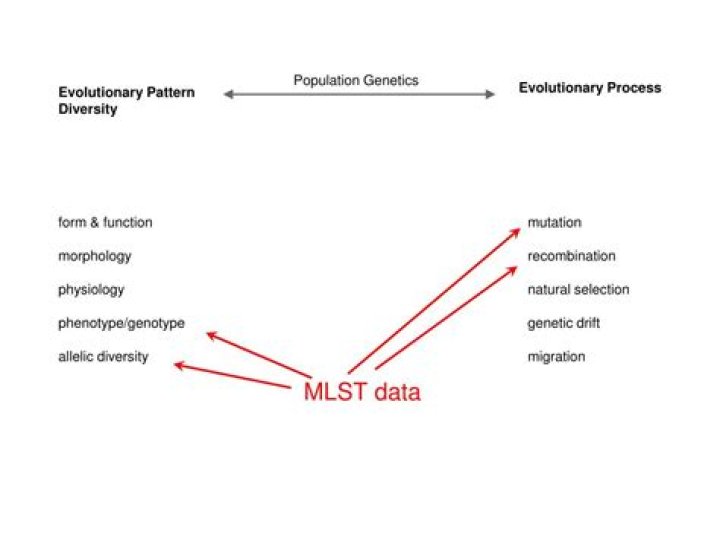
What is MLST database?
Multi-locus sequence typing (MLST) is a method of typing that facilitates the discrimination of microbial isolates by comparing the sequences of housekeeping gene fragments. The mlstdbNet software enables the implementation of distributed web-accessible MLST databases that can be linked widely over the Internet.
What is MLST used for?
Multilocus sequence typing (MLST) is an unambiguous procedure for characterising isolates of bacterial species using the sequences of internal fragments of (usually) seven house-keeping genes.
How is MLST performed?
The principle of MLST is simple: the technique involves PCR amplification followed by DNA sequencing. Nucleotide differences between strains can be checked at a variable number of genes depending on the degree of discrimination desired.
What is MLST analysis?
Multilocus sequence typing (MLST) is a molecular typing technique whereby a number of well chosen housekeeping genes (loci) are sequenced, usually in part. Instead, each sequence for a given locus is screened for identity with already known sequences for that locus. …
What class of genes is used in MLST analysis?
MLST was performed by DNA sequence analysis of seven housekeeping genes (aroE, ddl, dutA, tpi, recA, gmk, and sodA). The number of alleles ranged from five (dutA and ddl) to eleven (recA). Allelic profiles allowed the definition of 34 different sequence types (STs).
How do you cite PubMLST?
The preferred citation for this website is: Jolley et al. Wellcome Open Res 2018, 3:124 [version 1; referees: 2 approved]
What are the advantages of MLST?
One of the great advantages of MLST is its scalability from a single bacterial isolate to many hundreds or even thousands of samples. Upscaling of MLST is essential for large-scale studies and brings with it appreciable advantages in terms of reducing costs.
What is the difference between MLST and MLSA?
MLST was first proposed in 1998 by Maiden and colleagues. Moreover, MLST is usually applied to strains that belong to a well-defined species while MLSA is more often used when species boundaries are not well known and MLSA data are used to improve species descriptions.
What is BIGSdb?
BIGSdb is software designed to store and analyse sequence data for bacterial isolates. BIGSdb extends the principle of MLST to genomic data, where large numbers of loci can be defined, with alleles assigned by reference to sequence definition databases (which can also be set up with BIGSdb).
What is a clonal complex?
clonal complexes are defined as including any ST that matches the central genotype at four or more loci. If a particular strain has the same profile that defines a central allelic profile, then this ST number will at the same time both a clonal group and a clonal complex.
What is whole genome MLST?
Whole-genome MLST (wgMLST) analyses enable the recognition of genetic relationships between epidemiologically associated isolates and isolates that potentially have the same infection source.
What is PubMLST?
PubMLST – Public databases for molecular typing and microbial genome diversity.