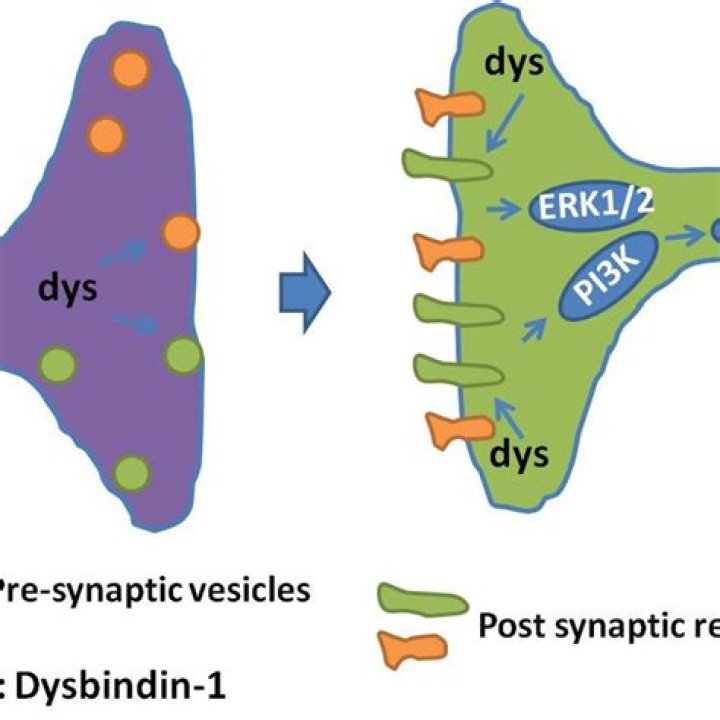
What is dysbindin 1?
Dysbindin-1 is a protein encoded by dystrobrevin-binding protein 1 gene (DTNBP1), which is located on the short (p) arm of chromosome 6 at position 22.3 [30]. Initially, dysbindin-1 was found to be a component of the dystrophin-associated protein complex (DPC) in skeletal muscle cells.
What is the difference between orthologs and paralogs?
“By definition, orthologs are genes that are related by vertical descent from a common ancestor and encode proteins with the same function in different species. By contrast, paralogs are homologous genes that have evolved by duplication and code for protein with similar, but not identical functions.”
Do orthologs have the same function?
Orthologs are defined as genes in different species that have evolved through speciation events only. It is generally assumed that orthologs have the same biological functions in different species [4], and duplication makes room for paralogs to evolve new functions [5].
Why do paralogs have different functions?
Once paralogs have been identified in a single genome, physical clustering by gene neighborhood can be used to group paralogs likely to have similar functions—because they physically group with the same genes across different genomes—and separate paralogs likely to have different functions—because they cluster with …
What is a paralogs?
Definition. Paralogous genes (or paralogs) are a particular class of homologous genes. They are the result of gene duplication and the gene copies resulting from the duplication are called paralogous of each other.
What do orthologs tell us?
Orthologs are genes in different species that evolved from a common ancestral gene by speciation, and, in general, orthologs retain the same function during the course of evolution. The display shows the maximum-likelihood phylogenetic tree representing the evolutionary history of gene families.
What is a orthologs?
Orthologs are defined as genes in different species that have evolved through speciation events only. A function-oriented ortholog group consists of orthologs that play the same biological role in different species and also includes recent paralogs with the same biological function, also known as “in-paralogs” [6].
What is homology in bioinformatics?
Homology is a concept that takes into account similarities that occur among nucleic acid or protein sequences of two different organisms. Homologous said to be orthologous if they were separated by an event called speciation.
Why are orthologs important?
Orthologs are defined as genes in different species that have evolved through speciation events only. Identification of orthologs accomplishes two goals: delineating the genealogy of genes to investigate the forces and mechanisms of evolutionary process, and creating groups of genes with the same biological functions.