How does T7 endonuclease work?
The T7 endonuclease 1 (T7E1) is a structure-selective enzyme that detects structural deformities in heteroduplexed DNA8. In using this assay to detect CRISPR-Cas9-mediated gene editing, reagents are transfected into cells, and the genomic DNA surrounding the target locus is amplified several days later by PCR.
How do I know if CRISPR is working?
How to Confirm Your CRISPR-cas9 Genome Editing Was Successful
- Check the Deletion.
- Sequence Your PCR Products.
- Measure Gene Expression.
- Measure Protein Expression.
- Measure the Impact in Your Cells or Model System.
- Share Your CRISPR Success with Anyone and Everyone!
What is mismatch cleavage assay?
A mismatch cleavage assay is a quick and easy way to detect indels. SurveyorTM nuclease is commonly used for this purpose, as it cleaves both DNA strands 3′ to any mismatches. It can detect indels of up to 12 nucleotides and is sensitive to mutations present at frequencies as low as 1 in 32 copies.
How is T7 endonuclease I different from a traditional restriction enzyme?
Endonuclease I is a junction-resolving enzyme encoded by bacteriophage T7, that selectively binds and cleaves four-way DNA junctions. However, it differs from a typical restriction enzyme as the proposed catalytic residues in both active sites are contributed by both polypeptides of the dimer.
What is crRNA CRISPR?
The gRNA is made up of two parts: crispr RNA (crRNA), a 17-20 nucleotide sequence complementary to the target DNA, and a tracr RNA, which serves as a binding scaffold for the Cas nuclease. The CRISPR-associated protein is a non-specific endonuclease.
Does CRISPR leave traces?
One important category of assisting technologies in CRISPR gene editing is methods used for detecting and quantifying indels (deletions or insertions). In addition, CRISPR-Cas9 can also leave footprints to the DNA without introducing DSBs, known as CRISPR’s DNA-binding footprints.
How do you confirm a knockout?
The 2 main ways to validate the knockout lines, will be firstly immunocytochemistry with the KO gene protein, and sequencing of the DNA to check whether or not it has been edited in the correct places.
How do you knock out a gene with CRISPR?
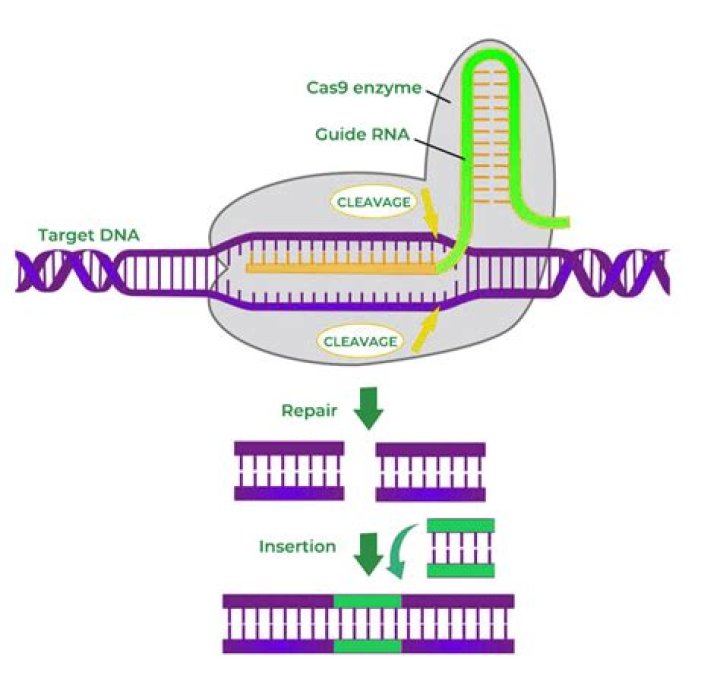
Knocking out a gene involves inserting CRISPR-Cas9 into a cell using a guide RNA that targets the tool to the gene of interest. There, Cas9 cuts the gene, snipping through both strands of DNA, and the cell’s regular DNA repair mechanism fixes the cut using a process called non-homologous end joining (NHEJ).
How do I change my restriction enzyme site?
To open the Destroy Restriction Site dialog, click Actions → Restriction Cloning → Destroy Restriction Site…, or simply press the Delete key.