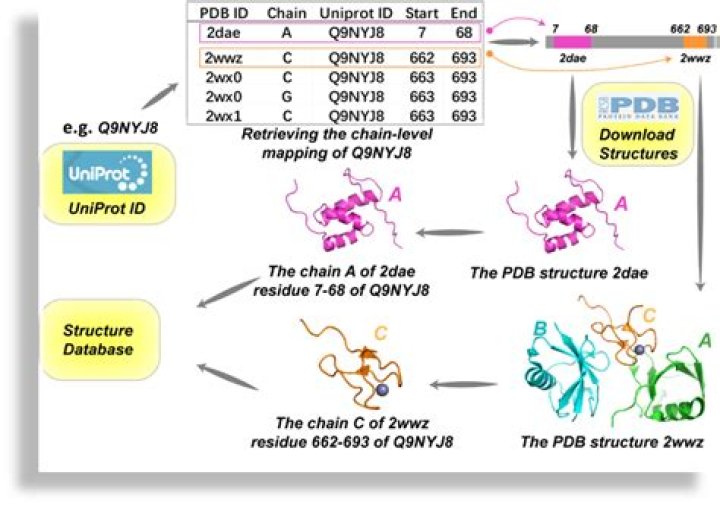
How do I download PDB from UniProt?
Using the query builder:
- Select Search in: Protein Knowledgebase (UniProtKB)
- Click Advanced Search » to open the query builder.
- Select Field: Cross-reference [DR]
- Select Database: PDB.
- Click Add & Search.
What is the UniProt database?
The Universal Protein Resource (UniProt) is a comprehensive resource for protein sequence and annotation data. UniProt is a collaboration between the European Bioinformatics Institute (EMBL-EBI), the SIB Swiss Institute of Bioinformatics and the Protein Information Resource (PIR).
How do I download proteome?
Go to the UniProt website and click on the search selection drop-down (Figure 60): Figure 60 Dataset selection drop-down. Select ‘Proteomes’, type Escherichia coli and click on the looking search icon (Figure 61): Figure 61 Proteomes search for ‘Escherichia Coli’.
Is UniProt a primary database?
UniProt is a freely accessible database of protein sequence and functional information, many entries being derived from genome sequencing projects….UniProt.
| Content | |
|---|---|
| Primary citation | UniProt Consortium |
| Access | |
| Data format | Custom flat file, FASTA, GFF, RDF, XML. |
| Website |
How do I download PDB files?
-Under Download the Structure File, right click on the X where the PDB(top) meets with none, under compression (on left) in the table and save target as. – Type in name of Protein / Macromolecule ( examples at bottom of page). – Click on Retrieve Released Data Matching Your Query icon.
What is UniProt and PDB?
PDB/UniProt Info retrieves annotations for Protein Data Bank (PDB) entries using a web service provided by the RCSB PDB. Sequences are displayed in Multalign Viewer, and feature annotations from UniProt are mapped onto the sequences as regions. See also: fetching UniProt data.
What is the difference between Swiss-Prot and UniProt?
UniProt provides a comprehensive, high-quality and freely accessible resource of protein sequence and functional information. UniProtKB/TrEMBL is a computer-annotated (unreviewed) supplement to Swiss-Prot, which strives to gather all protein sequences that are not yet represented in Swiss-Prot.
Is UniProt primary or secondary database?
Hybrid databases and families of databases Many data resources have both primary and secondary characteristics. For example, UniProt accepts primary sequences derived from peptide sequencing experiments. Some databases have different ‘branches’ for primary and secondary data.
How do I find my UniProt ID?
Select the Retrieve/ID mapping tab of the toolbar and enter or upload a list of identifiers (or gene names) to do one of the following: Retrieve the corresponding UniProt entries to download them or work with them on this website.
Is UniProt and Swiss Prot same?
UniProt provides a comprehensive, high-quality and freely accessible resource of protein sequence and functional information. UniProtKB/Swiss-Prot is a high quality manually annotated (reviewed) and non- redundant protein sequence database, which brings together experimental results and computed features.
Why do we use UniProt database?
UniProt helps with this in the following ways: It provides an up-to-date, comprehensive body of protein information at a single site. It aids scientific discovery by collecting, interpreting and organising this information so that it is easy to access and use. It provides tools to help with protein sequence analysis.